6.5 Types of Dominance
As we discussed in the previous section on polygenic traits, in humans most characteristics do not fit into two different phenotypes — complex traits, e.g., height, hair texture, skin colour etc., seemingly do not follow Mendelian analysis. As more scientists began analyzing genetic crosses using different types of plants and animals, it was found that while some traits obeyed Mendel’s laws (they were determined by a single gene with 1 dominant and 1 recessive allele), many other traits did not. In such cases, there were no definite recessive or dominant traits observed, or more than two alleles identified in a particular cross. In some instances, traits seem to be determined by more than one gene (multifactorial), and the environment also seemed to play a role through interaction with genes, to produce varying phenotypes.

These examples of the behaviour of certain traits implies a more complex array of interactions are at play, as these do not generate the typical Mendelian phenotypic ratios. We are extending Mendel’s Laws in order to provide explanations for the behaviour of such traits, and not necessarily challenging them.
One of the first concepts we need to understand, is that dominance is not always complete. Thus far, we have looked at the concept of dominance and recessiveness, whereby these conditions arise upon crossing two pure-breeding lines to create hybrids, and the hybrids are identical in phenotype to one parent for the particular trait in question. In this simplistic case, the allele passed down by that parent is said to be completely dominant when compared with the allele passed down by the parent whose trait is not manifested in the hybrid offspring. This type of arrangement is termed complete dominance.
As we will now see, there are two other types of Dominance — namely, incomplete dominance and co-dominance.
Complete Dominance
An example of a simple phenotype, is flower color in Mendel’s peas. We have already said that one allele as a homozygote produces purple flowers, while the other allele as a homozygote produces white flowers. But what about a heterozygous individual that has one purple allele and one white allele? What is the phenotype of a heterozygote?
This can only be determined by experimental observation. We know from observation that individuals heterozygous for the purple and white alleles of the flower colour gene have purple flowers. Thus, the allele associated with purple colour is, therefore, said to be dominant to the allele that produces the white colour. The white allele, whose phenotype is masked by the purple allele in a heterozygote, is recessive to the purple allele. The dominant/recessive character is a relationship between two alleles and must be determined by observation of the heterozygote phenotype.

Sometimes, to represent this relationship, a dominant allele will be written as a capital letter (e.g., A) while a recessive allele will be written in lower case (e.g., a). However, this is not the only system. Many different systems of genetic symbols are in use. The most common are shown in Table 6.5.1 Also note, genotypes (alleles) are usually written in italics and chromosomes and proteins are not. For example, the white gene in Drosophila melanogaster on the X chromosome encodes a protein called WHITE, which is a pigment precursor transmembrane transporter enzyme.
| Examples | Interpretation |
|---|---|
| A and a | Uppercase letters represent dominant alleles and lowercase letters indicate recessive alleles. Mendel invented this system but it is not commonly used because not all alleles show complete dominance and many genes have more than two alleles. |
| a+ and a1 | Superscripts or subscripts are used to indicate alleles. For wild type alleles the symbol is a superscript +. |
| AA or A/A | Sometimes a forward slash is used to indicate that the two symbols are alleles of the same gene locus, but on homologous chromosomes. |
Take a look at the video below. Incomplete Dominance, Codominance, Polygenic Traits, Epistasis, by Amoeba Sisters (2015) on YouTube, which discusses the various types of dominance and polygenic traits.
Incomplete Dominance
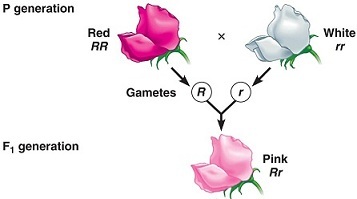
Other than the complete dominant and recessive relationship, other relationships can exist between alleles. In incomplete dominance (also called semi-dominance), both alleles affect the trait additively, and the phenotype of the heterozygote shows a typically intermediate between the homozygotes, which is often referred to as blended phenotype. For example, alleles for colour in carnation flowers (and many other species) exhibit incomplete dominance. Plants with alleles for red petals (RR) when crossed with a plant with alleles for white petals (rr) have offspring which have pink petals (Rr). We say that the R and the r alleles show incomplete dominance because neither allele is completely dominant over the other (Figure 6.5.3). Even though in Figure 6.5.3, there is the use of capital and common letters to indicate the two incompletely dominant alleles, a better way to represent such alleles would be the use of superscripts on the same letter e.g., R1 and R2.
Co-Dominance
Co-dominance is another type of allelic relationship in which a heterozygous individual expresses the phenotype of both alleles simultaneously. An example of co-dominance is found within the ABO blood group of humans. The ABO gene has three common alleles that were named (for historical reasons) IA, IB, and i. People homozygous for IA or IB display only A or B type antigens, respectively, on the surface of their blood cells, and therefore, have either type A or type B blood (Figure 6.5.4). Heterozygous IAIB individuals have both A and B antigens on their cells, and so have type AB blood. Note that the heterozygote expresses both alleles simultaneously, and is not some kind of novel intermediate between A and B. Co-dominance is, therefore, distinct from incomplete dominance, although they are sometimes confused.
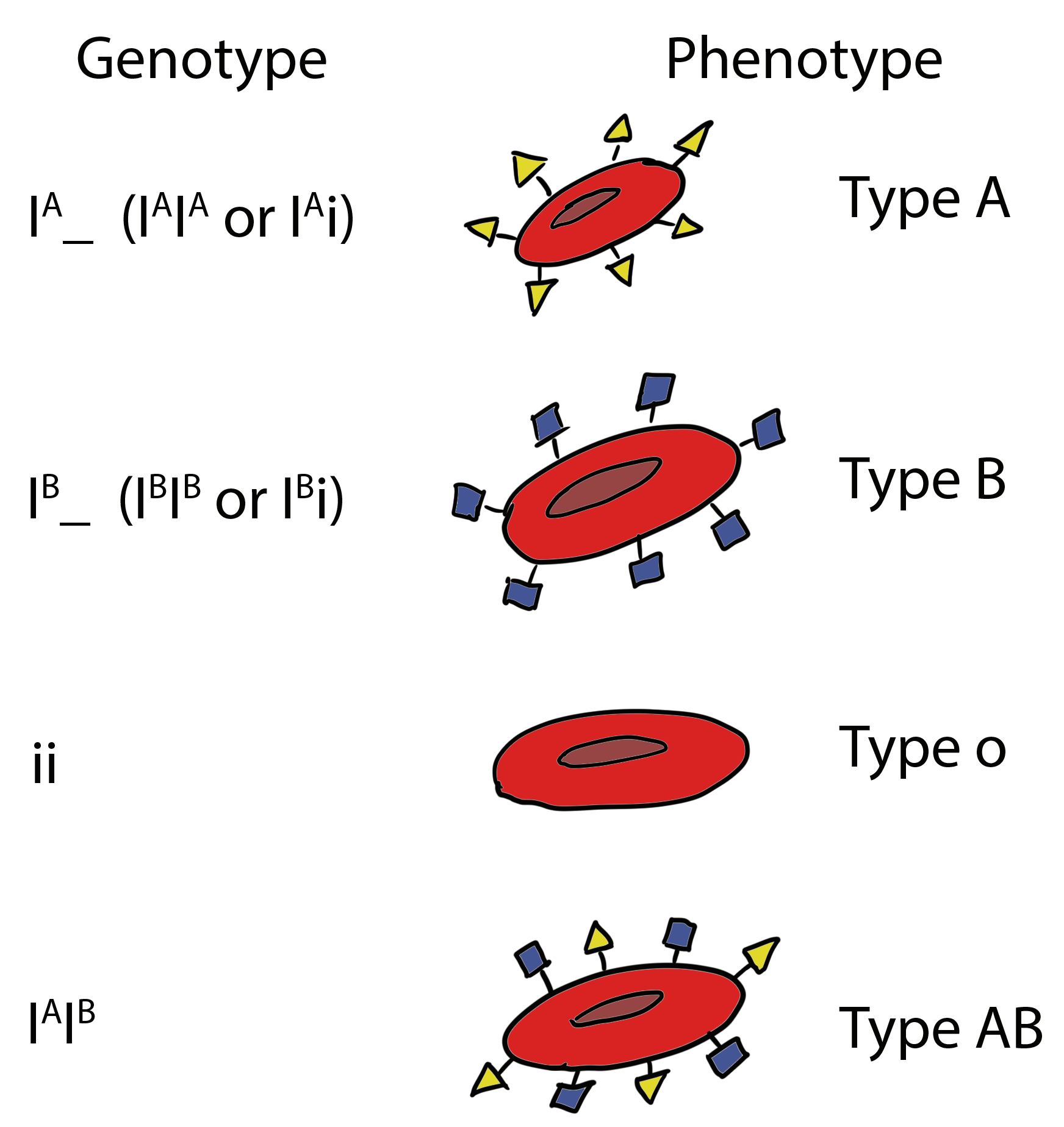
It is also important to note that the third allele, i, does not make either antigen and thus is recessive to the other alleles. IA/i or IB/i individuals display only A or B antigens, respectively. People homozygous for the i allele have type O blood.
This is a useful reminder that different types of dominance relationships can exist, even for alleles of the same gene.
Media Attributions
- Figure 6.5.1 2013 09 10 Tomate by Friedrich Haag, CC-BY-SA-4.0, via Wikimedia Commons
- Figure 6.5.2 14 05LawOfSegregation 2 L by Ashinkaaa, CC BY-SA 4.0, via Wikimedia Commons
- Figure 6.5.3 09 11aIncompleteDominance-L by RudLus02, CC BY-SA 4.0, via Wikimedia Commons
- Figure 6.5.4 Blood Type Codominance by DylanAudette, CC0 1.0 Universal Public Domain, via Wikimedia Commons
References
Amoeba Sisters. (2015, May 25). Incomplete dominance, codominance, polygenic traits, and epistasis! (video file). YouTube. https://www.youtube.com/watch?v=YJHGfbW55l0
Long Descriptions
- Figure 6.5.1 Two tomatoes: one is large and red in colour, and the other is small and orange in colour. The image seeks to demonstrate the variety of phenotypes that are possible with multifactorial traits, which are determined by more than one gene. [Back to Figure 6.5.1]
- Figure 6.5.2 Cross to demonstrate complete dominance. P generation is pure bred purple flowers crossed with pure breeding white flowers, resulting in all purple flowers (heterozygous) in the F1 generation. The F2 generation produces flowers in a ratio of 3 purple to 1 white, which is typical of complete dominance in a monohybrid cross. [Back to Figure 6.5.2]
- Figure 6.5.3 Cross to demonstrate incomplete dominance. P generation is pure-bred red flowers crossed with pure breeding white flowers. The F1 generation produced is comprised of pink flowers only, which is indicative of incomplete dominance between these two alleles controlling flower colour. [Back to Figure 6.5.3]
- Figure 6.5.4 The variety of blood types in humans. Four phenotypes are shown which are A, B, O and AB. These phenotypes are the result of combinations of alleles which exemplify co-dominance (A and B) as well as alleles which exemplify complete dominance (A and B over O). The combinations of alleles result on specific antigens being expressed on red blood cells of that organism, resulting in the four typical blood group phenotypes seen in humans. [Back to Figure 6.5.4]

