9.5 Unlinked Genes vs. Partial Linkage vs. Complete Linkage
When comparing any two genes, they can be varying distances apart. Their RF allows us to categorize them into the degree of linkage. The amount of linkage can be placed on a sliding scale.
Table 9.5.1 shows, generally, how we categorize the degree linkage using recombination frequency. Because RF is based upon experimental results that will have some experimental error, these should be treated as guidelines and not hard rules in determining the distance between genes.
| Linkage Description | Recombination Frequency |
|---|---|
| Unlinked | Approximately 50% or more than 35% |
| Partial linkage | More than 0% to 30% |
| Complete linkage | 0% |
Unlinked Genes
Unlinked genes appear to segregate and show independent assortment. There will be a random and even distribution of gamete types, and an RF of 0.50 is the expectation. This situation occurs in two instances: either when the genes are on completely different chromosomes, or when they are far enough apart on a single chromosome that crossovers are so numerous that alleles are distributed randomly (Figure 9.3.1). Either way, because the alleles are assorting independently, you should observe an equal number of recombinant and parental gametes, with an RF near ~0.50. Note, because of real-life variability, this value can be anywhere from ~0.40 to ~0.60.
Complete Linkage
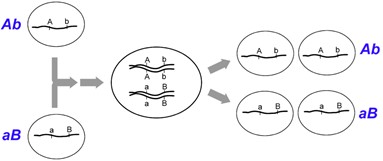
Having considered unlinked loci, let us turn to the opposite situation, in which two loci are so close together on a chromosome that the parental combinations of alleles always segregate together (Figure 9.5.1). This is because the physical distance between the two loci is so short that crossover events become extremely rare. Therefore, the alleles at the two loci are physically attached to the same chromatid and will nearly always segregate together into the same gamete. In this case, no recombinants will be present following meiosis, and the recombination frequency will be 0.00. This is complete (or absolute) linkage and is rare, as the loci must be so close together that crossovers are virtually impossible to detect.
Partial Linkage
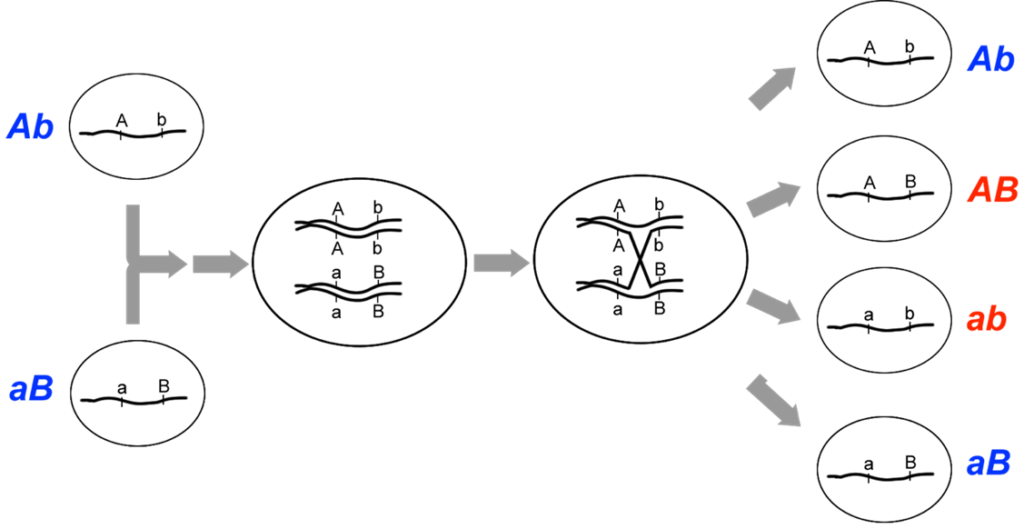
It is also possible to obtain recombination frequencies between 0% and 50%, which is a situation we call incomplete (or partial) linkage. Incomplete linkage occurs when two loci are located on the same chromosome, but the loci are far enough apart so that crossovers occur between them during some, but not all, meioses (Figure 9.5.2). Genes on the same chromosome are said to be syntenic regardless of whether they are completely or incompletely linked or unlinked. Thus, all linked genes are syntenic, but not all syntenic genes are linked.
Because the location of crossovers is essentially random for any given base pair of the chromosome, the greater the distance between two loci, the more likely a crossover will occur between them. Furthermore, loci on the same chromosome, but sufficiently separated from one another, will on average have multiple crossovers between them, and they will behave indistinguishably from physically unlinked loci. A recombination frequency of 50% is therefore the maximum recombination frequency that can be observed, and indicative of loci that are either on separate chromosomes, or sufficiently separated on the same chromosome.
Watch the video, (AP Biology) Linked Genes, Unlinked Genes, Incomplete Linkage, and Gene Mapping, by Mr. Cronin’s Videos (2019) on YouTube, which goes through a worked example involving linkage and gene mapping.
Media Attributions
- Figure 9.5.1 Original by Deyholos (2017), CC BY-NC 3.0, Open Genetics Lectures
- Figure 9.5.2 Original by Deyholos (2017), CC BY-NC 3.0, Open Genetics Lectures
References
Deyholos, M. (2017). Figures: 6. If two loci…; and 7. A crossover between …[digital image]. In Locke, J., Harrington, M., Canham, L. and Min Ku Kang (Eds.), Open Genetics Lectures, Fall 2017 (Chapter 18, p. 6). Dataverse/ BCcampus. http://solr.bccampus.ca:8001/bcc/file/7a7b00f9-fb56-4c49-81a9-cfa3ad80e6d8/1/OpenGeneticsLectures_Fall2017.pdf
Mr. Cronin’s Videos. (2019, December 18). (AP Biology) Linked genes, unlinked genes, incomplete linkage, and gene mapping (video file). YouTube. https://www.youtube.com/watch?v=J3AskTp1dsk
Long Descriptions
- Figure 9.5.1 Complete linkage of two loci results in gametes, where alleles will segregate in combinations identical to those present in the parental gametes (Ab, aB) and therefore no recombinants will be observed. [Back to Figure 9.5.1]
- Figure 9.5.2 The location of crossovers is essentially random for any given base pair of the chromosome; therefore, the greater the distance between two loci, the more likely a crossover will occur between them. A crossover between two linked loci can generate recombinant genotypes (AB ab), from the chromatids involved in the crossover. Multiple, independent meioses occur in each organism, so this particular pattern of recombination will not be observed among all the meioses from this individual. [Back to Figure 9.5.2]

