6.8 Muller’s Morphs
In this section, we will discuss the final class of mutants, called Muller’s Morphs. Most DNA sequence changes (mutations) occur at essentially random locations along a chromosome. Of those mutations occurring within genes, their mutant phenotypes (often recovered through genetic screens) are caused by loss-of-function alleles. These alleles are due to sequence changes in the DNA that cause a gene to produce fewer, less active, or non-active product (typically a protein), compared to the wild-type allele. Loss-of-function alleles tend to be recessive because the wildtype allele is haplo-sufficient. A loss-of-function allele that produces no active product is called an amorphs, or null, while alleles with only a partial loss-of-function are called hypomorphs.
More rarely, a mutant may have a gain-of-function allele, producing either more of the active product (hypermorphs) or producing an active product with a new and different function (neomorphs). Finally, antimorphs alleles have an activity that is dominant and opposite to the wild-type product’s function; antimorphs are also known as dominant negative mutations.
Thus, mutations (changes in a gene sequence) can result in mutant alleles that no longer produce the same level or type of active product as the wild-type allele. Any mutant allele can be classified into one of five types: (1) amorph, (2) hypomorph, (3) hypermorph, (4) neomorph, and (5) antimorph.
Amorph
Amorphic alleles have a complete loss-of-function. They make no active product — zero function. They are known as a “Null” mutation or a “loss-of-function” mutation.
Molecular explanation: Changes in the DNA base pair sequence of an amorphic allele may cause one or more of the following:
- Gene deletion — The DNA sequence is removed from the chromosome.
- Gene is present, but is not transcribed because of a sequence change in the promoter or enhancer/regulatory elements.
- Gene is present, but the transcript is aberrantly processed. There is normal transcription but base pair changes cause the mature mRNA to incorrectly splice introns, therefore the translated amino acid sequence would be altered and nonfunctional.
- Gene is present and a transcript is produced but no translation occurs — changes in the base pair sequences would preclude the mRNA from binding to the ribosome for translation.
- Gene is present and a transcript is produced and translated but a nonfunctional protein product is produced — the mutation alters a key amino acid in the polypeptide sequence producing a completely non-functional polypeptide.
Genetic/phenotypic explanation: Amorphic mutations of most genes usually act as recessive to wild type (Case #1). However, with some genes the amorphic mutations are dominant to wild type (Case #2).
Cases 1 & 2
case #1: white gene in Drosophila
| Allele Combination | Result |
|---|---|
| w+/w+ | wildtype and red eyed |
| w+/w– | wildtype and red eyed |
| w–/w– | mutant and white eyed |
case #2: Minute locus in Drosophila
| Allele Combination | Result |
|---|---|
| M+/M+ | wildtype and long bristled |
| M+/M– | mutant and short bristled |
| M–/M– | dead, recessive lethal |
For the Minute gene, we concluded that the organism needs both copies to have a wild type phenotype. Loss of one copy (an amorphic mutation) produces a dominant visible mutant phenotype. Deletion of the gene is an example of a classic amorphic mutation.
Hypomorph
Hypomorphic alleles show only a partial loss-of-function. These alleles are sometimes referred to as “leaky” mutations, because they provide some function, but not complete, normal function.
Molecular explanation: Changes in the DNA base pair sequence of the hypomorphic allele may cause one or more of the following, with the gene still present.
- Reduced transcription — changed DNA sequence in the promoter or enhancer/regulatory elements can reduce the level of transcription.
- Aberrant processing of the transcript — normal transcription but base pair changes cause the mature mRNA to incorrectly splice introns, therefore the translated protein sequence would be altered and function at a reduced level.
- Reduced translation — changes in the base pair sequences would reduce the efficiency of the mRNA binding to the ribosome for translation.
- Reduced-function protein product — normal transcription, processing, and translation, but mutation changes certain amino acid in the polypeptide sequence so its function is reduced.
Genetic/phenotypic explanation: Hypomorphic mutations of most genes usually act as recessive to wild type, though hypomorphic mutations theoretically could be dominant to wildtype.
Hypomorph
white-apricot allele in Drosophilia
| Allele Combination | Result |
|---|---|
| w+/w+ | wildtype ad red eyed |
| w+/wa | wildtype and red eyed |
| wa/wa | mutant and apricot eye colour |
Both amorphs and hypomorphs tend to be recessive to wild type in diploids because the wild type allele is usually able to supply sufficient product to produce a wild type phenotype (called haplo-sufficient). If the mutant allele is not able to produce a wild type phenotype, it is haplo-insufficient, and it will be dominant to the wild type allele. Here -/+ heterozygotes produce a mutant phenotype.
While the first two classes involve a loss-of-function, the next two involve a gain-of-function — quantity or quality. Gain-of-function alleles are almost always dominant to the wild type allele.
Hypermorph
Hypermorphic alleles produce quantitatively more of the same, active product.
Molecular explanation: Changes in the DNA base pair sequence of the hypermorphic allele may cause one or more of the following, with the gene still present.
- Increased transcription — changed DNA sequence in the promoter or enhancer/regulatory elements that increase the level of transcription.
- Increased translation — changes in the base pair sequences would increase the efficiency of the mRNA binding to the ribosome for translation.
- Increased function protein product — normal transcription, processing, translation, but base pair changes alter certain amino acids in the polypeptide sequence, so its function is normal but increased in amount.
Genetic/phenotypic explanation: Hypermorphic mutations of most genes usually act as dominant to wild type since they are a gain of function, The classic hypermorph is a gene duplication.
Neomorph
Neomorphic alleles produce a product with a new, different function, something that the wild type allele does not do.
Molecular explanation: Changes in the DNA base pair sequence of the neomorphic allele may cause one or more of the following, with the gene still being present.
- New transcription — changed DNA sequence in the promoter or enhancer/regulatory elements that makes new transcription either temporally or in a tissue-specific manner.
- New function protein product — normal transcription, processing, translation but base pair changes alter certain amino acids in the polypeptide sequence so it acquires a new function (activity) that is different from the normal function (e.g., additional substrate or new binding site).
Genetic/phenotypic explanation: Most neomorphic mutations act dominant to wild type since they are a gain-of-function. The classical neomorphic mutation is a translocation that moves a new regulatory element next to a gene promoter so it is expressed in a new tissue or at a new time during development. Such mutations are often produced when chromosome breaks are rejoined and the regulatory sequences of one gene are juxtaposed next to the transcriptional unit of another, creating a novel, chimeric gene.
Antimorph
Antimorphic alleles are relatively rare, and have a new activity that is dominant and opposite to the wildtype function. These alleles usually interfere with the function from the wild type allele. (They often lose their normal function as well.) The new function works against the normal expression of the wild type allele. This can happen at the transcriptional, translational, or later level of expression. Thus, when an antimorphic allele is heterozygous with wild type, the wild type allele function is reduced or prevented. At the molecular level, there are many ways this can happen. The simplest model to explain an antimorphic effect is that the protein acts as a dimer (or any multimer) and the inclusion of a mutant subunit poisons the whole complex, thereby preventing or reducing its level of function. Antimorphs are also known as dominant-negative mutations because they are usually dominant and act negatively against the wild type function.
Identifying Muller’s Morphs
All mutations can be sorted into one of the five morphs based on how they behave with heterozygous with three other standard alleles (Figure 6.8.1): (1) deletion alleles (zero function), (2) wild type alleles (normal function), and (3) duplication alleles (double normal function).
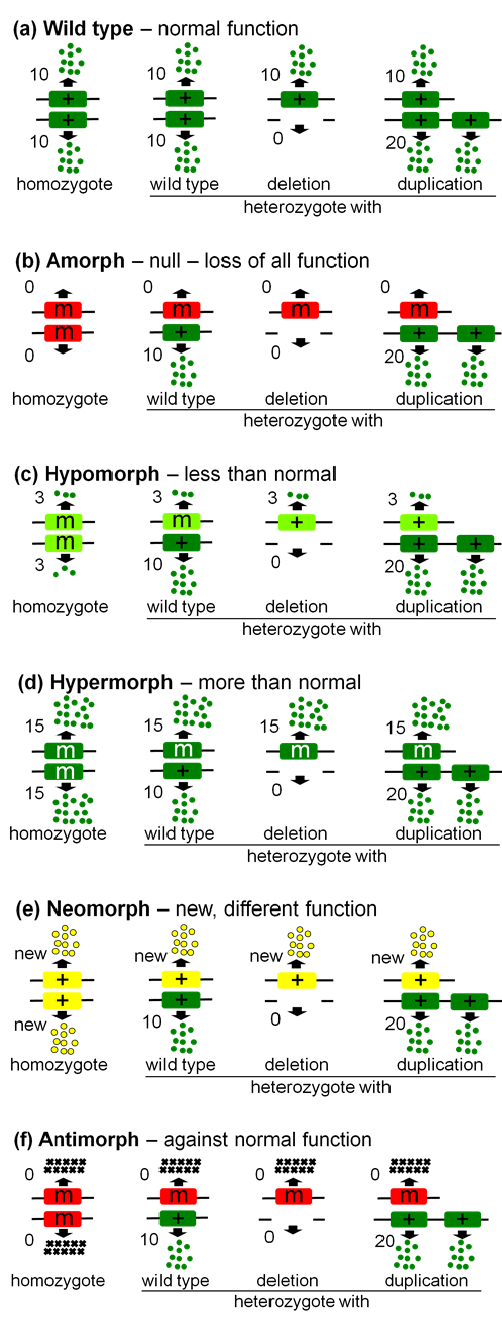
Loss-of-Function
- Haplo-insufficient — a loss-of-function dominant mutation
- Null mutations — loss-of-function mutation that remove the activity of the gene product
- Hypomorphic mutations — decreased activity of the gene product — loss-of-function mutant phenotypes can be due to partial or complete elimination of the activity of a gene’s encoded product
Gain-of-function
- Hypermorph — mutation that produces more gene activity per gene dose than wild-type
- Neomorph — mutation that produces some novel gene activity that is not characteristic of the wild-type
Media Attributions
- Figure 6.8.1 Original by Locke (2017), CC BY-NC 3.0, Open Genetics Lectures
References
Locke, J. (2017). Figure 7. Five classes of mutants designated as morphs [5 digital images]. In Locke, J., Harrington, M., Canham, L. and Min Ku Kang (Eds.), Open Genetics Lectures, Fall 2017 (Chapter 13, p. 8). Dataverse/ BCcampus. http://solr.bccampus.ca:8001/bcc/file/7a7b00f9-fb56-4c49-81a9-cfa3ad80e6d8/1/OpenGeneticsLectures_Fall2017.pdf
Long Description
- Figure 6.8.1 (a-f) Comparison of the wild type expresses normal function with the five types of mutants known as Muller’s Morphs. The Amorph shows a loss of all function; the hypomorph shows less than normal function; the hypermorph shows more than normal function; the neomorph shows a new different function, and the antimorph produces a product which works against normal function. [Back to Figure 6.8.1]

